Identification and Functional Characterization of microRNAs in Ichthyosporea
Zhu, H. and Bråte, J. 2025. F1000Research. doi:https://doi.org/10.12688/f1000research.162501.1
Abstract
Background
microRNAs (miRNAs) are key post-transcriptional regulators of gene expression in animals, yet their origins and early functions remain elusive. In this study, we explore the diversity and functionality of miRNAs in Ichthyosporea, a group of unicellular organisms that represent some of the closest relatives of animals.
Methods
We have performed miRNA identification by integrating deep small RNA sequencing with draft genome assemblies across six species of Ichthyosporea, including Abeoforma whisleri, Pirum gemmata, and Sphaeroforma sp. To assess the functional roles of these miRNAs, we have used computational target prediction and degradome sequencing.
Results
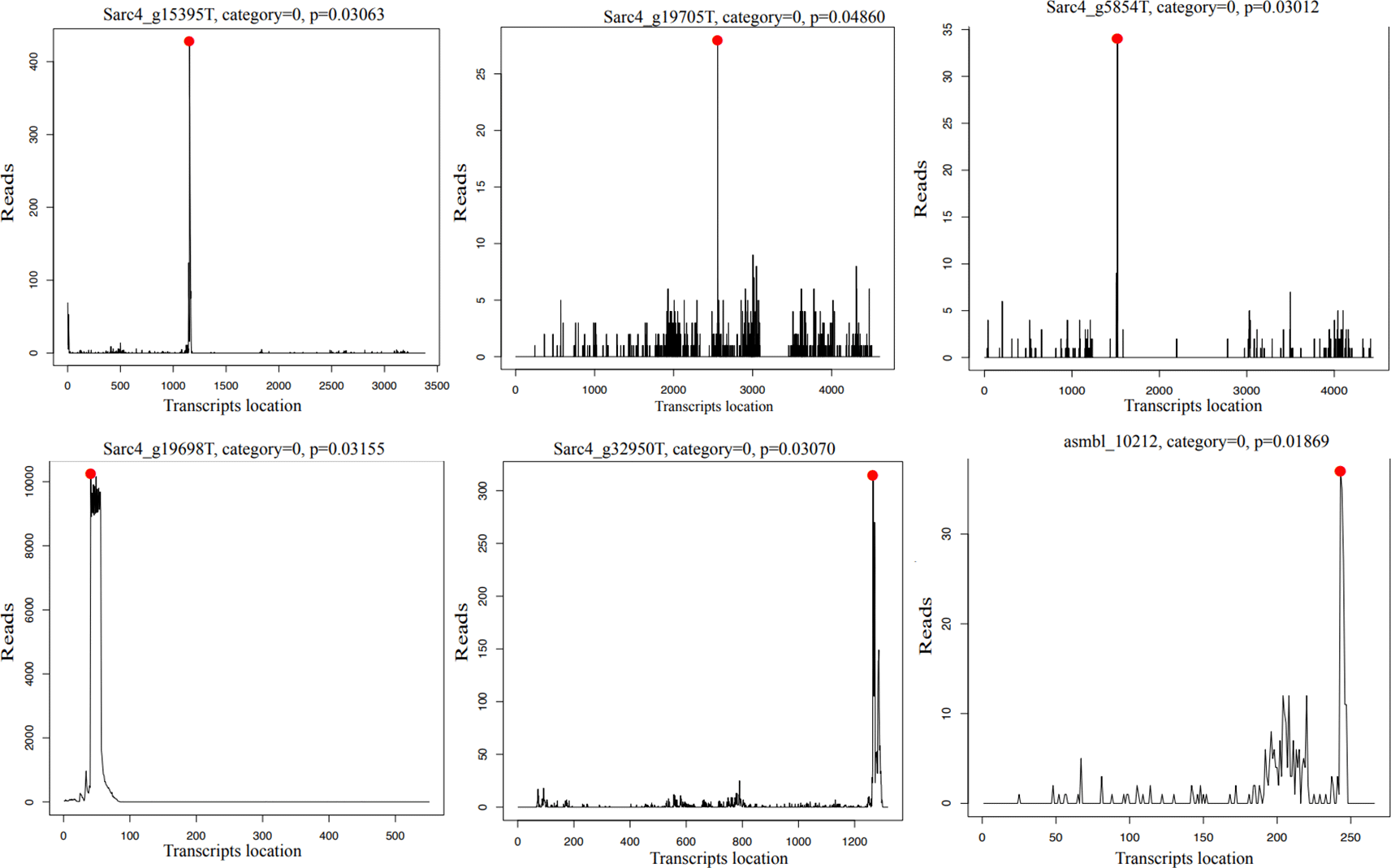
We identified a rich and varied repertoire of miRNAs across Ichthyosporea. Many of these miRNAs appear to be lineage-specific, while a subset is conserved among closely related species, suggesting both rapid evolutionary turnover and the persistence of ancient regulatory elements. Moreover, predicted target genes span a wide range of transcripts, implicating miRNAs in diverse cellular processes. Additionally, degradome sequencing reveal that in S. arctica, several miRNAs are likely to induce mRNA cleavage.
Conclusions
These findings not only extend previous reports of miRNA presence in Ichthyosporea but also underscore the deep evolutionary roots of miRNA-mediated gene regulation in the holozoan lineage. The miRNA induced target cleavage in S. arctica is also observed in cnidarians and supports the hypothesis that mRNA cleavage may represent the ancestral function of animal miRNAs.